
Bioinformatics is growing in the clinical field, and in my job in a clinical microbiology lab, I’m increasingly asked for tips about how to get into bioinformatics or genomic data science.
As I recently took the plunge into genomics from my PhD field of small molecular structures and machine learning, as part of …

Ad-hoc tasks in bioinformatics can contain such an immense number of operations and tasks that need to be performed to achieve a certain goal. Often these are all individually regarded as rather “standard” or “routine”. Despite this, it is quite hard to find an authoritative set of …

As we are - according to some expert opinions - living in the Century of Biology, I found it interesting to reflect on Go’s usage within the field.
Go has some great features that make it really well suited for biology, such as:
A relatively simple language that can be learned in a short time even for people …
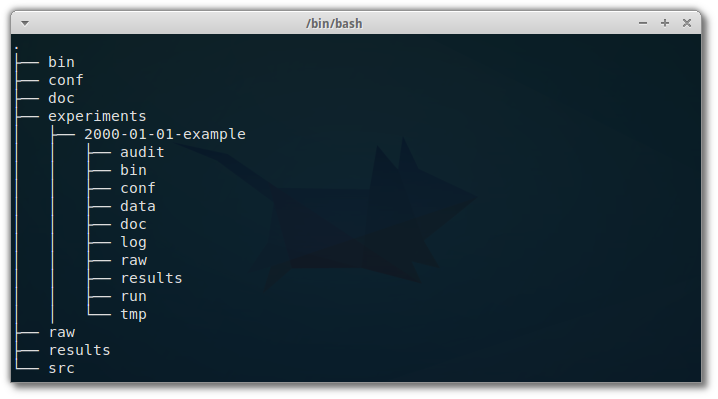
I read this excellent article with practical recommendations on how to organize a computational project, in terms of directory structure.
Directory structure matters The importance of a good directory structure seems to often be overlooked in teaching about computational biology, but can be the difference between a …

It turned out I didn’t have the time and strength to blog every day at the NGS Bioinformatics Intro course, so here comes a wrap up with some random notes and tidbits from the last days, including any concluding remarks!
These days we started working on a more realistic NGS pipeline, on analysing re-sequencing …

Today was the second day of the introductory course in NGS bioinformatics that I’m taking as part of my PhD studies.
For me it started with a substantial oversleep, probably due to a combination of an annoying cold and the ~2 hour commute from south Stockholm to Uppsala and BMC . Thus I missed some really …

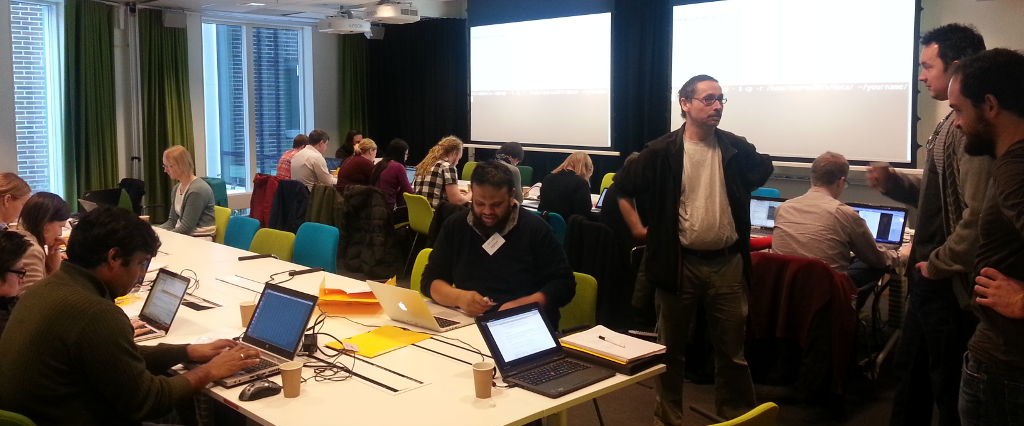
Just finished day 1 of the introductory course on Bioinformatics for Next generation sequencing data at Scilifelab Uppsala. Attaching a photo from one of the hands-on tutorial sessions, with the tutorial leaders, standing to the right.
Today’s content was mostly introductions to the linux commandline in general, …
Right now I’m sitting on the train and trying to get my head around some of the pre-course materials .






